Do you know the chip used for nucleic acid detection?
Nucleic acid detection is essentially realized by using the PCR reaction (polymerase chain reaction) we learned in high school biology books. The PCR reaction utilizes the double-strand replication principle of DNA to amplify a large number of target genes in vitro. The nucleic acid amplification stage requires the use of a PCR instrument, which can automatically "denature-anneal-extend" the submitted liquid to amplify the amount of nucleic acid. It seems very simple, but in fact it is very time-consuming from nucleic acid collection to results.
The collected sample, the swab, is kept in a collection tube filled with red fluid. The red liquid is the virus preservation solution, which contains ingredients that can sterilize and inactivate the virus and a moderate pH value, which can not only inactivate the virus but also protect the nucleic acid of the virus from being damaged by the outside world. As for why it is red, it is actually convenient for subsequent staff to perform pipetting operations, and it is also convenient for distinguishing different detection types. Transshipment is to transport the red test tubes one by one to the testing institution for subsequent operations. If they cannot be transshipped in time, they need to be stored at low temperature.
Since RNA is single-stranded, before nucleic acid detection, it is necessary to reverse transcribe the RNA of the virus into a double-stranded DNA, and then amplify it by the PCR technique mentioned above to achieve the necessary nucleic acid concentration for detection. Then use the method of fluorescent group labeling to detect whether the sample contains the accounting sequence to be tested, that is, to detect whether it contains the gene fragment unique to the new coronavirus. The PCR instrument can monitor that the number of cycles (Ct value) at which the fluorescence reaches a preset threshold is related to the concentration of viral nucleic acid. The higher the concentration of viral nucleic acid, the smaller the Ct value.

After the test is completed, the staff will pay attention to uploading the positive or negative results uniformly, and then you will see the nucleic acid test results on your mobile phone.
We focus on the process of nucleic acid detection. After information verification and registration, the staff first need to transfer the liquid in the test tube that may contain viral nucleic acid fragments to another test tube with a pipette gun, and use special reagents or spin column methods (simply put, the virus in the virus Nucleic acid is thrown out) for nucleic acid extraction, and then the nucleic acid value is added by the PCR technology mentioned above. Among them, the process of pipetting and PCR value-added operations are the most time-consuming steps, and this is also the longest process for nucleic acid detection except for external factors.
The timeliness of nucleic acid detection is very important, so automated and portable detection has become the key to improving the overall detection speed, and our protagonist today - microfluidic chip came into being.
A microfluidic chip that balances efficiency and accuracy
Microfluidics (Microfluidics) is a technology and science that uses micron-scale flow channels to process and process micro-volumes (1 nanoliter to 1 Attoliter, 1 nanoliter = 0.000001 ml) of liquid. The flow channel in the microfluidic chip (Microfluidic Chip), on the one hand, can be used as a reaction vessel for biochemical related experiments. Under the control of components such as valves and pumps, complex biochemical processes can be controlled automatically and on a large scale. On the other hand, at the microscale, the fluid in the microfluidic chip will present a predictable standard laminar flow, which makes the modeling of the system and the prediction of reaction kinetics easier, making microfluidics evolve into an ideal A medium for studying chemical reactions. Due to its great potential in the fields of biology, chemistry, and medicine, it has developed into a new research field interdisciplinary in biology, chemistry, medicine, fluid, electronics, materials, and machinery.
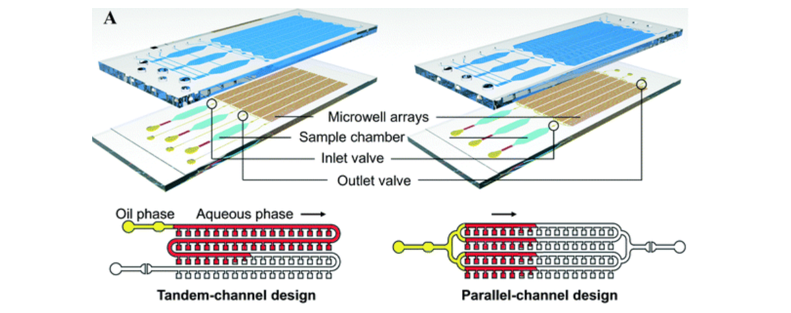
Back to microfluidic chips and nucleic acid detection. The combination of microfluidic chip and PCR technology can effectively reduce the volume of the reaction device. Multiple microfluidic chips can also be placed in a single PCR instrument to react at the same time, reducing the overall reaction time and completing nucleic acid amplification in a shorter time. ; In addition, less reaction solution can also effectively improve the detection sensitivity; microfluidics also has a certain degree of portability due to the reduced volume, which can respond to sudden outbreaks faster.
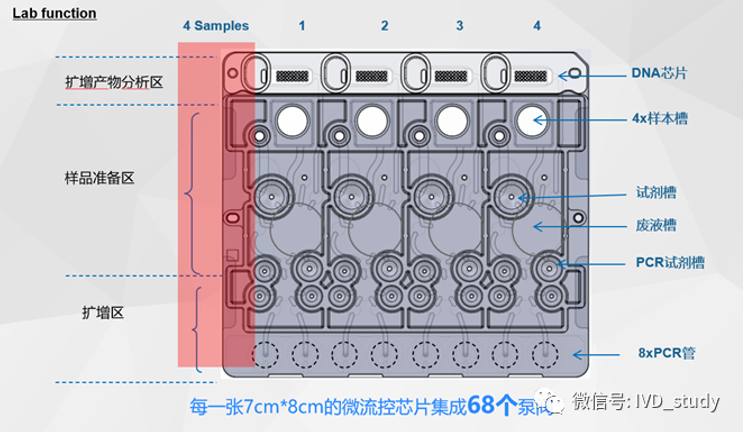
So with the PCR technology supported by microfluidic chips, how much can its computing power be improved?
Microfluidic chips are actually not chips in the traditional sense. It has no working voltage, no memory, or even electricity, and naturally has no parameters such as computing power and threads. The microfluidic chip, why is it a chip?


